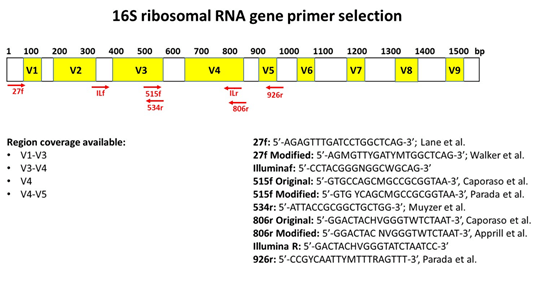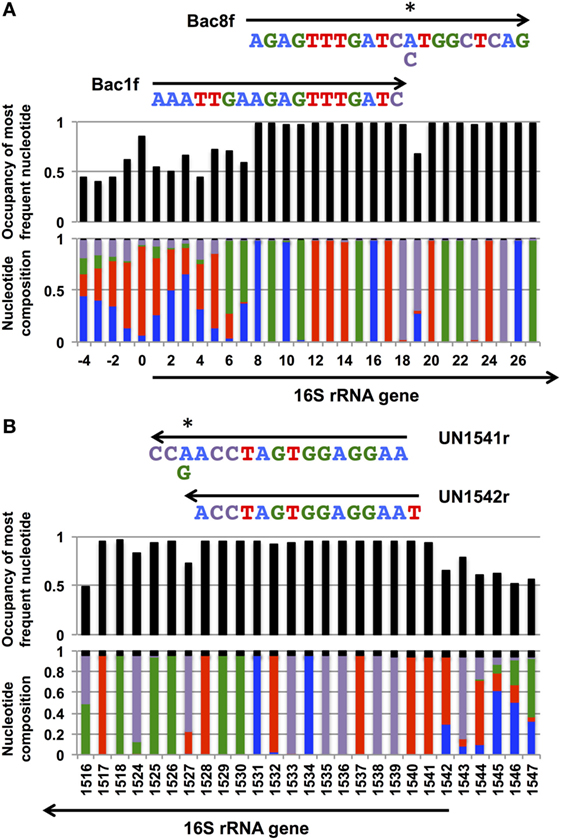
Frontiers | PCR Primer Design for 16S rRNAs for Experimental Horizontal Gene Transfer Test in Escherichia coli

16S rRNA Gene Primer Validation for Bacterial Diversity Analysis of Vegetable Products - ScienceDirect

Absolute quantitation of microbes using 16S rRNA gene metabarcoding: A rapid normalization of relative abundances by quantitative PCR targeting a 16S rRNA gene spike‐in standard - Zemb - 2020 - MicrobiologyOpen - Wiley Online Library
Development of a Prokaryotic Universal Primer for Simultaneous Analysis of Bacteria and Archaea Using Next-Generation Sequencing | PLOS ONE

PCR primers used for amplification of 16S rRNA gene for cyanobacteria... | Download Scientific Diagram
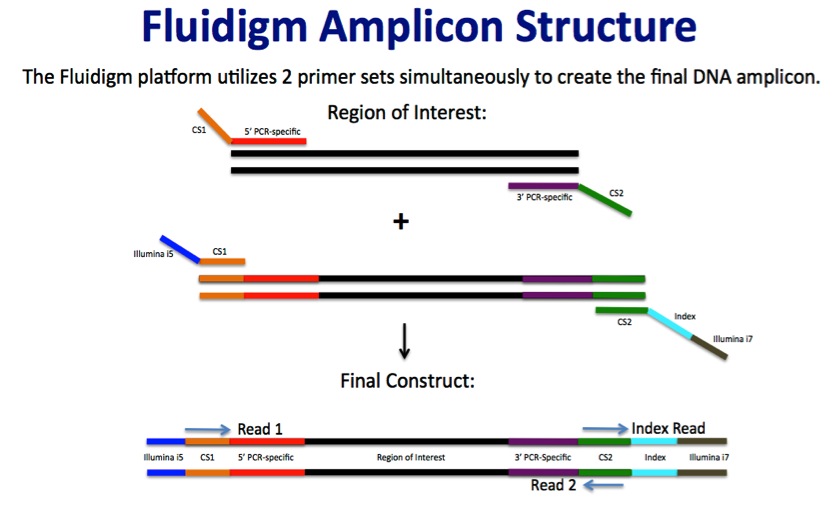
HTDNA: Services/Equipment: Preparation of 16S rRNA (and all other) Amplicons for Illumina Sequencing | Biotech

Improved Multiplex PCR Using Conserved and Species-Specific 16S rRNA Gene Primers for Simultaneous Detection ofActinobacillus actinomycetemcomitans, Bacteroides forsythus, and Porphyromonas gingivalis | Journal of Clinical Microbiology

16S rRNA gene primer choice impacts off-target amplification in human gastrointestinal tract biopsies and microbiome profiling | Scientific Reports

Table 1 from Design and evaluation of universal 16S rRNA gene primers for high-throughput sequencing to simultaneously detect DAMO microbes and anammox bacteria. | Semantic Scholar

Sequences of the forward (F) and reverse (R) primers, for 16S rRNA gene | Download Scientific Diagram

Better skin microbiome analyses using new 16S V4 region primers developed by Microbiome Insights' scientific team — Microbiome Insights